d3viz – d3viz: Interactive visualization of Aesara compute graphs#
Guide#
Requirements#
d3viz requires the pydot
package. pydot-ng fork is better
maintained, and it works both in Python 2.x and 3.x. Install it with pip:
pip install pydot-ng
Like Aesara’s printing module, d3viz
requires graphviz binary to be available.
Overview#
d3viz extends Aesara’s printing module to interactively visualize compute
graphs. Instead of creating a static picture, it creates an HTML file, which can
be opened with current web-browsers. d3viz allows
to zoom to different regions and to move graphs via drag and drop,
to position nodes both manually and automatically,
to retrieve additional information about nodes and edges such as their data type or definition in the source code,
to edit node labels,
to visualizing profiling information, and
to explore nested graphs such as OpFromGraph nodes.
Note
This userguide is also available as
IPython notebook.
As an example, consider the following multilayer perceptron with one hidden layer and a softmax output layer.
import aesara as th
import aesara.tensor as at
import numpy as np
ninputs = 1000
nfeatures = 100
noutputs = 10
nhiddens = 50
rng = np.random.RandomState(0)
x = at.dmatrix('x')
wh = th.shared(rng.normal(0, 1, (nfeatures, nhiddens)), borrow=True)
bh = th.shared(np.zeros(nhiddens), borrow=True)
h = at.sigmoid(at.dot(x, wh) + bh)
wy = th.shared(rng.normal(0, 1, (nhiddens, noutputs)))
by = th.shared(np.zeros(noutputs), borrow=True)
y = at.special.softmax(at.dot(h, wy) + by)
predict = th.function([x], y)
The function predict outputs the probability of 10 classes. You can
visualize it with aesara.printing.pydotprint() as follows:
from aesara.printing import pydotprint
import os
if not os.path.exists('examples'):
os.makedirs('examples')
pydotprint(predict, 'examples/mlp.png')
The output file is available at examples/mlp.png
from IPython.display import Image
Image('./examples/mlp.png', width='80%')
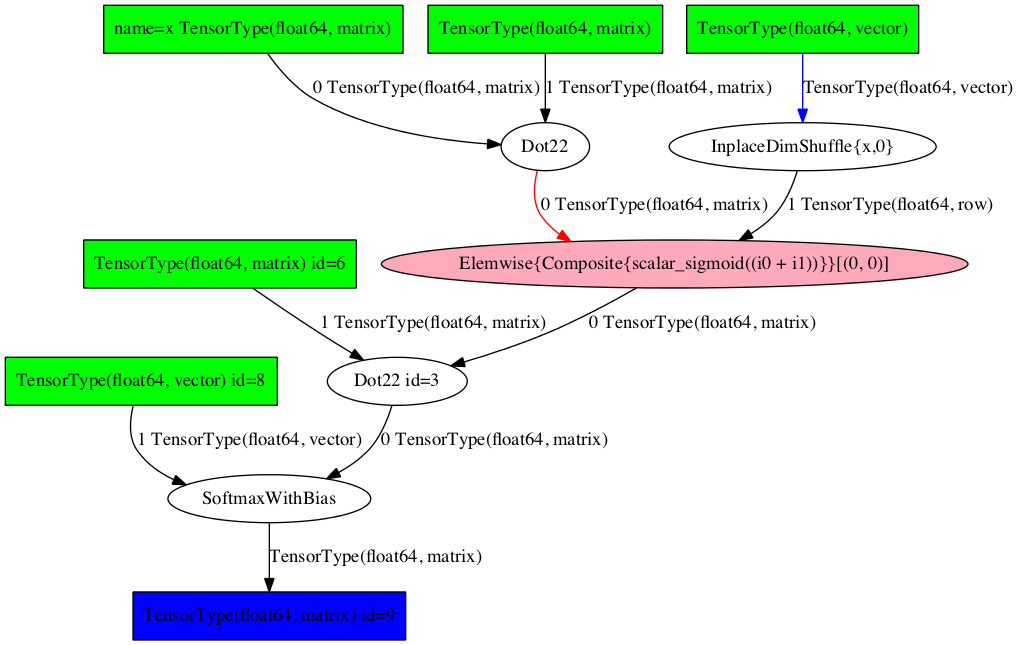
To visualize it interactively, import aesara.d3viz.d3viz.d3viz() from
the the aesara.d3viz.d3viz module, which can be called as before:
import aesara.d3viz as d3v
d3v.d3viz(predict, 'examples/mlp.html')
When you open the output file mlp.html in your web-browser, you will
see an interactive visualization of the compute graph. You can move the
whole graph or single nodes via drag and drop, and zoom via the mouse
wheel. When you move the mouse cursor over a node, a window will pop up
that displays detailed information about the node, such as its data type
or definition in the source code. When you left-click on a node and
select Edit, you can change the predefined node label. If you are
dealing with a complex graph with many nodes, the default node layout
may not be perfect. In this case, you can press the Release node
button in the top-left corner to automatically arrange nodes. To reset
nodes to their default position, press the Reset nodes button.
You can also display the interactive graph inline in
IPython using IPython.display.IFrame:
from IPython.display import IFrame
d3v.d3viz(predict, 'examples/mlp.html')
IFrame('examples/mlp.html', width=700, height=500)
Currently if you use display.IFrame you still have to create a file, and this file can’t be outside notebooks root (e.g. usually it can’t be in /tmp/).
Profiling#
Aesara allows function profiling via the profile=True flag. After at least
one function call, the compute time of each node can be printed in text form
with debugprint. However, analyzing complex graphs in this way can be
cumbersome.
d3viz can visualize the same timing information graphically, and
hence help to spot bottlenecks in the compute graph more easily! To
begin with, we will redefine the predict function, this time by
using profile=True flag. Afterwards, we capture the runtime on
random data:
predict_profiled = th.function([x], y, profile=True)
x_val = rng.normal(0, 1, (ninputs, nfeatures))
y_val = predict_profiled(x_val)
d3v.d3viz(predict_profiled, 'examples/mlp2.html')
When you open the HTML file in your browser, you will find an additional
Toggle profile colors button in the menu bar. By clicking on it,
nodes will be colored by their compute time, where red corresponds to a
high compute time. You can read out the exact timing information of a
node by moving the cursor over it.
Different output formats#
Internally, d3viz represents a compute graph in the Graphviz DOT
language, using the
pydot package, and defines a
front-end based on the d3.js library to visualize
it. However, any other Graphviz front-end can be used, which allows to
export graphs to different formats.
formatter = d3v.formatting.PyDotFormatter()
pydot_graph = formatter(predict_profiled)
pydot_graph.write_png('examples/mlp2.png');
pydot_graph.write_png('examples/mlp2.pdf');
Image('./examples/mlp2.png')
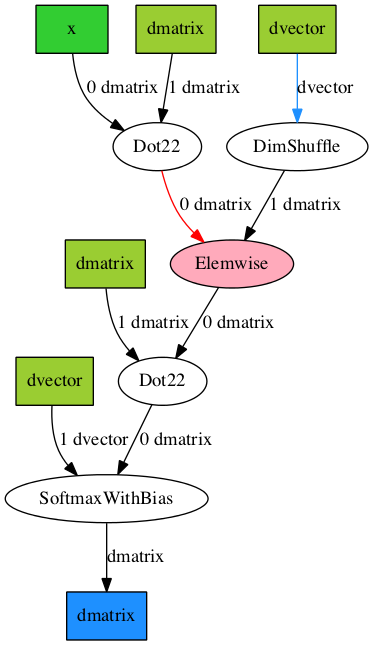
Here, we used the aesara.d3viz.formatting.PyDotFormatter class to
convert the compute graph into a pydot graph, and created a
PNG and PDF
file. You can find all output formats supported by Graphviz here.
OpFromGraph nodes#
An OpFromGraph node defines a new operation, which can be called with
different inputs at different places in the compute graph. Each OpFromGraph
node defines a nested graph, which will be visualized accordingly by d3viz.
x, y, z = at.scalars('xyz')
e = at.sigmoid((x + y + z)**2)
op = th.compile.builders.OpFromGraph([x, y, z], [e])
e2 = op(x, y, z) + op(z, y, x)
f = th.function([x, y, z], e2)
d3v.d3viz(f, 'examples/ofg.html')
In this example, an operation with three inputs is defined, which is used to build a function that calls this operations twice, each time with different input arguments.
In the d3viz visualization, you will find two OpFromGraph nodes,
which correspond to the two OpFromGraph calls. When you double click on
one of them, the nested graph appears with the correct mapping of its
input arguments. You can move it around by drag and drop in the shaded
area, and close it again by double-click.
An OpFromGraph operation can be composed of further OpFromGraph operations, which will be visualized as nested graphs as you can see in the following example.
x, y, z = at.scalars('xyz')
e = x * y
op = th.compile.builders.OpFromGraph([x, y], [e])
e2 = op(x, y) + z
op2 = th.compile.builders.OpFromGraph([x, y, z], [e2])
e3 = op2(x, y, z) + z
f = th.function([x, y, z], [e3])
d3v.d3viz(f, 'examples/ofg2.html')
Feedback#
If you have any problems or great ideas on how to improve d3viz,
please let me know!
Christof Angermueller
References#
d3viz module#
Dynamic visualization of Aesara graphs.
Author: Christof Angermueller <cangermueller@gmail.com>
- aesara.d3viz.d3viz.d3viz(fct, outfile, copy_deps=True, *args, **kwargs)[source]#
Create HTML file with dynamic visualizing of an Aesara function graph.
In the HTML file, the whole graph or single nodes can be moved by drag and drop. Zooming is possible via the mouse wheel. Detailed information about nodes and edges are displayed via mouse-over events. Node labels can be edited by selecting Edit from the context menu.
Input nodes are colored in green, output nodes in blue. Apply nodes are ellipses, and colored depending on the type of operation they perform.
Edges are black by default. If a node returns a view of an input, the input edge will be blue. If it returns a destroyed input, the edge will be red.
- Parameters:
fct (aesara.compile.function.types.Function) – A compiled Aesara function, variable, apply or a list of variables.
outfile (str) – Path to output HTML file.
copy_deps (bool, optional) – Copy javascript and CSS dependencies to output directory.
Notes
This function accepts extra parameters which will be forwarded to
aesara.d3viz.formatting.PyDotFormatter.
- aesara.d3viz.d3viz.d3write(fct, path, *args, **kwargs)[source]#
Convert Aesara graph to pydot graph and write to dot file.
- Parameters:
fct (aesara.compile.function.types.Function) – A compiled Aesara function, variable, apply or a list of variables.
path (str) – Path to output file
Notes
This function accepts extra parameters which will be forwarded to
aesara.d3viz.formatting.PyDotFormatter.
PyDotFormatter#
- class aesara.d3viz.formatting.PyDotFormatter(compact=True)[source]#
Create
pydotgraph object from Aesara function.- Parameters:
compact (bool) – if True, will remove intermediate variables without name.
- __call__(fct, graph=None)[source]#
Create pydot graph from function.
- Parameters:
fct (aesara.compile.function.types.Function) – A compiled Aesara function, variable, apply or a list of variables.
graph (pydot.Dot) –
pydotgraph to which nodes are added. Creates new one if undefined.
- Returns:
Pydot graph of
fct- Return type:
pydot.Dot
